CRIMSON
(Click on images for larger version)
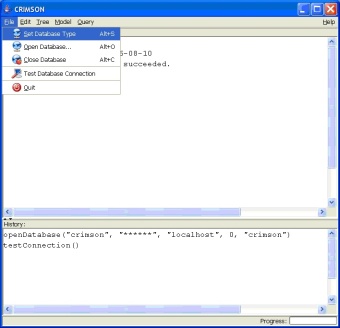
GUI Menu Panel: The GUI has a main window which contains various menus, a "Messages" window and a "History" window. The 'Messages' window will display all status, warning, and error messages. The 'History' window contains the command line equivalents for all commands initiated via the GUI.
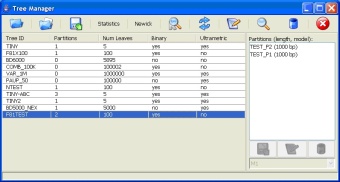
Tree Manager: The tree manager facilitates loading new trees, adding or removing data partitions, deleting existing trees, adding notes to trees, etc.
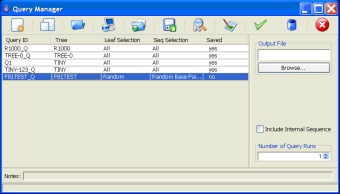
Query Manager: A graphical query manager allows users to manipulate existing queries and add new queries. When users create queries they have the option of publishing the queries to the database so that they are available in future sessions and to other users. It is also possible to export queries to text files so that they can be shared between databases.
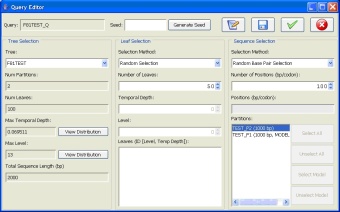
Query Editor: Users are provided with graphical means to run existing queries and to easily set the parameters associated with the running of queries. For example, a user can specify that a query run multiple times to repeatedly sample from a tree using the same query specifications.
Model Editor: Crimson stores information about the data models used and can link data models to trees and data partitions.
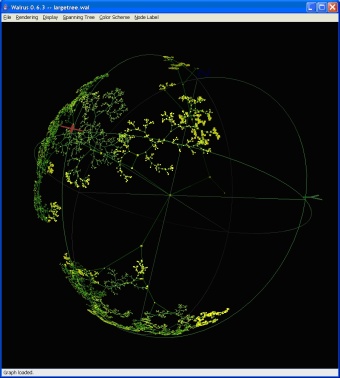
3D Tree Viewer: To facilitate the viewing of trees, Crimson contains routines to easily display trees in the 3D tree viewer Walrus.